During early mammalian development, the totipotent zygote undergoes the first cell fate decision to form a blastocyst that includes inner cell mass (ICM) and trophectoderm (TE). ICM will give rise to epiblast, which generates all future embryonic lineages, and primitive endoderm (PrE), which forms future yolk sac. By contrast, trophectoderm primarily differentiates to the placenta that supports the development of embryos. These two cell fates are supported by two classes of lineage transcription factors (TFs). Evidence from in vitro stem cells indicate that these two classes of regulators typically promote their respective programs, but inhibit the opposing programs. Intriguingly, before the first cell fate determination, many ICM regulators and TE regulators already co-exist within the same cells. Do the lineage TFs prime their controlled cell fate programs at this stage, and inhibit the opposing programs? How do early embryos resolve the conflict from the co-expression of these factors and their potential predisposition of two cell fates? Due to the scarcity of early embryo research materials, these questions remain as unresolved mysteries in the field.
On January 19, 2024, the research group led by Professor Wei Xie from the School of Life Sciences at Tsinghua University published a research article in the journal Nature Structural & Molecular Biology, entitled " Lineage regulators TFAP2C and NR5A2 function as bipotency activators in totipotent embryos". This study revealed that lineage specifying TFs play distinct roles in the initiation and commitment stages of the first cell fate decision in mammalian embryos. The researchers found two lineage TFs, TFAP2C and NR5A2, activate both ICM and TE programs in mammalian totipotent embryos, leading to ‘bipotency activation’ to initiate the first cell fate specification. At later stages upon lineage commitment, these lineage TFs gradually transit to unidirectional activators, through extensive genome-wide chromatin re-localization.
TFAP2C and NR5A2, which are generally thought to be TE and pluripotency regulators, respectively, are both highly expressed at the 8-cell stage. The researchers first investigated the chromatin targets of TFAP2C via CUT&RUN during the early mouse lineage specification across 6 stages, spanning the initiation, commitment, and maintenance phases. Moreover, they also generated Tfap2c knockout embryos to dissect its functions. Unexpectedly, at the initiation stage, TFAP2C bound and activated both early TE and ICM genes at the bipotent stages, rather than specifically activating the TE fate program. These results indicate lineage TFs do not prime cell fate programs, instead function as “bipotency activators”.
The researchers further investigated how TFAP2C becomes TE-promoting factor at later stages. In TE cells, TFAP2C leaves early ICM genes but retains its binding to early TE genes. It further binds and activates late TE genes, ultimately achieving specific activation of the TE program. The exact mechanism by which TFAP2C accomplishes this transition is not entirely clear, and the researchers speculate that it may be associated with the activation of other TE lineage-specific factors. For instance, CDX2 is another key TE regulator expressed later than TFAP2C. The researchers observed substantial co-localization of TFAP2C and CDX2 binding in TE cells, and knockout of TFAP2C could partially impact CDX2 binding. This cooperation between transcription factors can even occur between different lineage factors. In the post-implantation extra-embryonic ectoderm (ExE), researchers found TFAP2C co-localizing with the classical pluripotency factor SOX2 and regulating the expression of extra-embryonic genes. The loss of TFAP2C also affected the chromatin binding of SOX2.
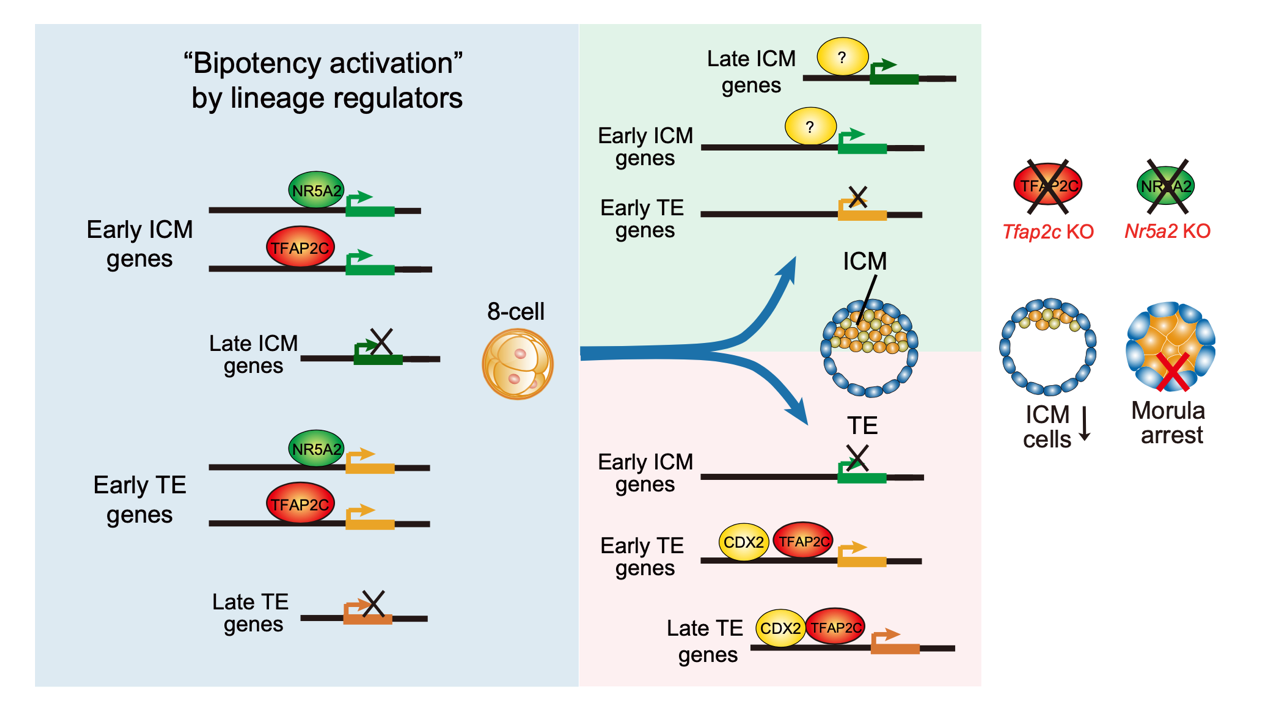
A model for bipotency activation by lineage regulators during the first cell fate specification
Furthermore, such “bipotency activation” in totipotent embryos also applied to NR5A2. NR5A2 bound and activated both ICM and TE lineage genes in 8-cell embryos, then predominantly regulated the ICM program in ESCs.
Based on these data, the researchers propose that TFs should not be simply defined as lineage factors. This is because the functions of TFs in different cells may be entirely different (for example, SOX2 can promote embryonic lineage programs in the epiblast but can also promote extra-embryonic programs in ExE). Rather, multiple TFs can interact to form a lineage module, which may be key to determine a cell identity. The functions of TFs can be more accurately defined defined only when considered its related lineage modules. Together, these data not only reveal a unique transcription circuity in early embryos, but also give insights for the investigation of TFs in different cell contexts.
Prof. Wei Xie is the corresponding author of this paper. Ph.D. students Lijia Li and Fangnong Lai from the School of Life Sciences at Tsinghua University are co-first authors. The research associate Ling Liu, postdoctoral Xukun Lu, Ph.D. student Xiaoyu Hu, and postdoctoral Bofeng Liu from Prof. Wei Xie's research group also made important contributions to this study. This study also received help from the Animal Center and the Bioinformatics core facility at Tsinghua University. Funding for this research was provided by the National Natural Science Foundation of China, the Key Research and Development Program of the Ministry of Science and Technology, and the Tsinghua-Beijing Life Science Center. Prof. Wei Xie is an HHMI International Research Scholar and a New Cornerstone investigator.
Article link: https://www.nature.com/articles/s41594-023-01199-x
