During the long-term fighting against pathogens, plants have evolved a sophisticated innate immune system to defend against infection. Plants pattern recognition receptors (PRRs) sense pathogen-associated molecular patterns (PAMPs) or host-derived danger-associated molecular patterns (DAMPs) to activate an immune response termed pattern-triggered immunity (PTI). PRRs are mainly composed of receptor kinases (RKs) and receptor-like proteins (RLPs). Specifically, one subgroup of PRRs contains an extracellular leucine-rich repeat (LRR) domain called LRR-type PRR, which can be further divided into LRR-RKs and LRR-RLPs. Although the molecular mechanism of ligand recognition and activation of LRR-RKs is well understood, however, mechanisms underlying ligand recognition and activation of LRR-RLPs, which play a critical role in plant immunity, remain elusive.
On September 21 of 2022, Jijie Chai's group from Tsinghua University and Yuanchao Wang's group from Nanjing Agricultural University published a paper entitled "Plant receptor-like protein activation by a microbial glycoside hydrolase" in Nature. This study reveals a conserved mechanism of ligand-induced heterodimerization of an LRR-RLP with BAK1 and suggests a dual function of the LRR-RLP in plant immunity.
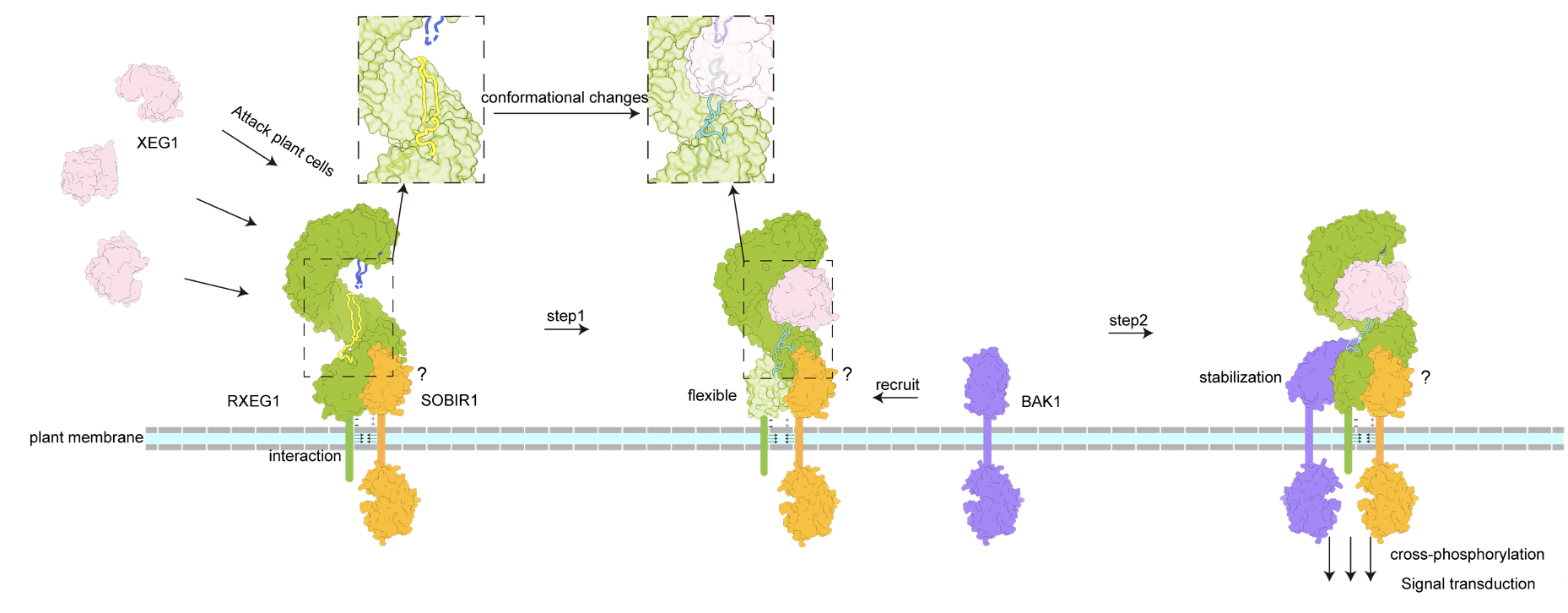
Figure 1 Working model of XEG1-induced RXEG1 activation.
The research groups led by Dr. Chai and Dr. Wang found that RXEG1 binds to XEG1 mainly through two unique loops formed by the extracellular N-terminal cap and C-terminal island region. Comparison of the crystal structure of XEG1-RXEG1 and cryo-EM structure of apo-RXEG1 showed that XEG1 binding causes significant conformational changes in the RXEG1 island region and C-terminus, leading to the formation of the XEG1-RXEG1-BAK1 complex. The last four LRRs disordered in the XEG1-RXEG1 structure become well defined upon BAK1 binding. The structures together with biochemical data unraveled the mechanisms of ligand recognition by an LRR-RLP and ligand-induced association of the LRR-RLP with its co-receptor BAK1, providing a structural paradigm for understanding the activation mechanism of LRR-RLPs.
Interestingly, the two loop regions of RXEG1 bind to the active site pocket of XEG1. Previous studies by Yuanchao Wang's group have shown that XEG1’s xyloglucanase enzyme activity is essential for Phytophthora virulence. The in-vitro enzymatic assay and in-vivo function study also proved that RXEG1 inhibit XEG1’s xyloglucanase activity. Taken together, data from this study revealed for the first time the molecular mechanism of ligand recognition and activation of plant LRR-RLPs, and identified a dual function of plant RLPs immunity. The study provides a paradigm for understanding the plant RLP family.

Figure 2 Structure of RXEG1-XEG1-BAK1 complex
Prof. Jijie Chai from School of Life Science, Tsinghua University, and Prof. Yuanchao Wang from School of Plant Protection, Nanjing Agricultural University, Associate professor Zhifu Han from School of Life Science, Tsinghua University, and Associate professor Yan Wang from School of Plant Protection, Nanjing Agricultural University are co-corresponding authors. Yue Sun, graduate students from the School of Life Sciences of Tsinghua University, Yan Wang, an associate professor at the School of Plant Protection, Nanjing Agricultural University, and Xiaoxiao Zhang, a former postdoctoral fellow in the School of Life Sciences of Tsinghua University and currently an associate professor at Shanghai University of Science and Technology, were co-first authors of the paper.
The National Center for Protein Science (Beijing), the cryo-electron microscope platform of Tsinghua University, the high-performance computing platform of Tsinghua University and the Shanghai Synchrotron Radiation Light Source provided technical support for this research.
The research was supported by the National Natural Science Foundation of China, the National Soybean Industry Technology System Project, the National Key Research and the Alexander von Humboldt Foundation, the Max-Planck-Gesellschaft, Deutsche Forschungsgemeinschaft.
https://www.nature.com/articles/s41586-022-05214-x
