Prof. Jijie Chai’s group at School of Life Sciences, Tsinghua University, in collaboration with Prof. Jane Parker’s and Prof. Paul Schulze-Lefert’s groups from the Max Planck Institute for Plant Breeding Research, reported on the molecular mechanisms by which the plant TNL disease resistance protein RPP1 recognizes the effector protein ATR1 and assembles into an ATR1-induced resistosome to form a holoenzyme for NAD+ hydrolysis. The study provides a paradigm for understanding the activation mechanism of plant TNLs.
This study was published online in Science on December 4th, 2020, entitled “Direct pathogen-induced Assembly of an NLR immune receptor Complex to Form a Holoenzyme”. At the same time, a perspective on the paper was also published in Science in the same issue.
Jijie Chai's group has been working on the mechanisms of action of NLR family receptors. Following the previous report of the auto-inhibition and indirect activation mechanisms of the CNL ZAR1, this study revealed the molecular mechanism of direct ligand recognition by the TNL RPP1 and activation of the enzymatic activity of the receptor.


Perception of pathogen- or host-derived components by immune receptors is essential for plant survival and initiates powerful defense responses to resist the invasion of pathogenic microorganisms. Nucleotide binding domain (NBD) and leucine-rich repeat (LRR) containing receptors (NLR) are a major class of plant immune receptors which specifically recognize pathogen effectors to induce effector-triggered-immunity (ETI). According to their N-terminal domains, plant NLRs are mainly divided into two classes: CNLs with an N-terminal coiled-coil domain (CC) and TNLs with an N-terminal toll/interleukin receptor (TIR). Both classes of receptor proteins can recognize pathogenic effector proteins in a direct or indirect manner. Studies from Chai’s group last year found that the CNL ZAR1 from Arabidopsis thaliana can form a large protein complex termed resistosome upon indirect recognition of an effector protein. The ZAR1 resistosome may function as an ion channel or a transporter for effector-triggered immunity. Compared to CNLs, the immune signaling mechanism of TNLs appears more complicated. The N-terminal TIR domain of TNLs has NAD+hydrolase activity (NADase), which is supposed to generate second messenger molecules activating an EDS1 and NRG1-dependent immune pathway. Many questions remainded about TNL signaling. For example, how do TNLs recognize pathogenic effector proteins? Can TNLs oligomerize to form a ZAR1-like resistosome upon activation? Do full-length TNLs possess NADase activity similar to that of the TIR domain? If they do, how does recognition of effector proteins regulate NADase activity of TNLs?
Arabidopsis RPP1 is a canonical TNL, which is composed of an N-terminal TIR domain, followed by a NOD (nucleotide binding and oligomerization domain) and a C-terminal LRR domain. ATR1 is an effector of the pathogen Hyaloperonospora arabidopsidis and its direct recognition by RPP1 triggers ETI.
Formation of the RPP1 resistosome induced by ATR1
In this study, the authors co-expressed RPP1 and ATR1 in insect cells and purified the RPP1-ATR1 complex termed the RPP1 resistosome using a tandem purification procedure. They then resolved a structure of the complex by cryo-EM to a resolution of 3.16 Å (Fig. 1).
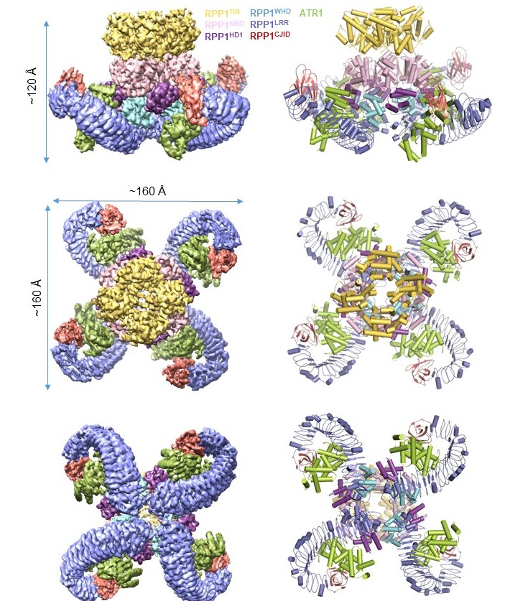
Fig. 1 Tetrameric structure of the RPP1 resistosome
Structural basis of direct ATR1 recognition by RPP1
The Cryo-EM structure shows that the LRR domain of RPP1 directly engages in ATR1 binding. Importantly, structural comparison revealed a novel domain at the C-terminus, designated as C-terminal Jelly roll and Ig-like domain(C-JID), which also participates in the direct binding of ATR1. Further analysis shows that the C-JID is present in almost all TNLs, suggesting that this domain may have a broad role in TNL ligand recognition. ATR1 binding induces conformational changes in RPP1, relieving intramolecular auto-inhibition of RPP1 and consequently assembling into a tetrameric complex termed the RPP1 resistosome (Fig. 1). This structure provides strong evidence for direct recognition of effector proteins by NLRs.
The RPP1 resistosome functions as a holoenzyme for NAD+ hydrolysis
Biochemical analyses showed that formation of the RPP1 resistosome greatly promoted its NAD+ hydrolysis activity, which was further enhanced by divalent metal ions such as magnesium and calcium ions. These results indicate that the RPP1 resistosome functions as a holoenzyme to catalyze NAD+ hydrolysis. The cryo-EM structure provides significant insights into the mechanism underlying the RPP1 resistosome as a holoenzyme enzyme. In contrast to the other structural domains of the receptor with C4 symmetry, the four TIR domains in the RPP1 resistosome are arranged with C2 symmetry. This results in the formation of two head-to-tail TIR homodimers, which interact with each other to form the C2 symmetry-related tetramers. Multiple lines of evidence support the head-to-tail TIR homodimers function as a complete NADase catalyzing NAD+ hydrolysis. First, an ATP molecule that may mimic NAD+ binds to a head-to-tail TIR homodimer; second, a structural comparison showed that the ATP molecule has a similar binding site to NADP+, another substrate of the TIR domain of RUN1; finally, mutations of key amino acids from the BB-loop critical for the formation of the head-to-tail TIR homodimers significantly reduced the NAD+ hydrolysis activity of the RPP1 resistosome and ATR1-induced ETI.
The RPP1 resistosome is ADP-bound
It is generally believed that ligand binding promotes exchange of ADP (bound to the NOD module) in the inhibited state with ATP/dATP, resulting in activation and oligomerization of NLRs. Intriguingly, the cryo-EM structure shows that the NOD of the RPP1 resistosome binds an ADP instead of ATP/dATP as seen in the ZAR1 resistosome. ZAR1 uses the arginine residue (R) from the highly conserved ‘TTR’ motif of CNLs to coordinate the γ-phosphate group of ATP/dATP, thus stabilizing the active conformation of ZAR1. However, the residue corresponding to the arginine residue from the TTR motif of ZAR1 is substituted with a glutamic acid residue in RPP1, thus disrupting ATP/dATP binding activity of the TNL.
Graduate student Shoucai Ma from School of Life Science at Tsinghua University, postdoctoral fellows Dmitry Lapin and Li Liu from Max Planck Institute for Plant Breeding, Germany; graduate student Yue Sun from School of Life Science at Tsinghua University and postdoctoral fellow Wen Song from University of Cologne, are co-first authors of this paper. Prof. Jijie Chai, from the School of Life Science at Tsinghua University, Prof. Jane Parker and Prof. Paul Schulze-Lefert from the Max Planck Institute for Plant Breeding Research are co-corresponding authors. Former postdoctoral fellow Xiao Zhang from Beijing Advanced Innovation Center for Structural Biology, Elke Logemann and Dongli Yu from the Max Planck Institute for Plant Breeding Research, Jan Jirschitzka from the University of Cologne, and associate-investigators Wang Jia and Han Zhifu from the School of Life Science at Tsinghua University are also involved in the study. This study was supported by the National Natural Science Foundation of China, Beijing Advanced Innovation Center for Structural Biology, Tsinghua-Peking Joint Center for Life Sciences, the Alexander von Humboldt Foundation and German Research Foundation.
Paperlink: https://science.sciencemag.org/content/370/6521/eabe3069
