Prof. Wei Xie’ group at Tsinghua University has published an article in Nature Genetics, entitled ‘Epigenomic analysis of gastrulation identifies a unique chromatin state for primed pluripotency’ on December 17, 2019. By employing multiple ultra-sensitive chromatin analyses, they revealed a unique chromatin state of primed pluripotency in mouse post-implantation embryos, including unusually strong enrichment of H3K4me3/H3K27me3 (super bivalency) at key developmental promoters. Their findings not only uncovered lineage-specific chromatin reconfiguration during early cell fate commitment, but also revealed how epigenetic transition occurs from naive pluripotency to primed pluripotency in vivo.
Around implantation, epiblast cells transit from na?ve to primed pluripotency, and further give rise to three germ layers, including ectoderm, mesoderm and endoderm. This provides the foundation of the entire body development. Yet, how chromatin reconfigures during this critical developmental window in vivo remains elusive. By integrating genome-wide investigation of histone modifications, chromatin accessibility, and 3D chromatin architecture during this period, the researchers discovered that enhancers in ectoderm, but not for endoderm or mesoderm, are already pre-accessible in E6.5 epiblast, supporting the notion that ectoderm is the default fate for epiblast. Interestingly, they found that unusually strong H3K4me3 emerges at developmental gene promoters specifically in E6.5 epiblast and E7.5 ectoderm, but not for earlier stages and committed lineages. Together with H3K27me3, these promoters form super bivalency. Furthermore, super bivalency marked developmental genes showed strong spatial interactions specifically in E6.5 epiblast. To investigate how super bivalency is established and regulated, the researchers knocked out Kmt2b, which encoded a methyltransferase specifically catalyzing bivalent H3K4me3, in zygote. They found that deficiency of KMT2B leads to loss of H3K4me3, reduced spatial clustering in E6.5 epiblast and impaired subsequent activation for a subset of developmental genes, ultimately resulting in embryonic lethality. Thus, their study provided comprehensive spatiotemporal views for the dynamic chromatin architecture in mouse post-implantation embryos, and further revealed super bivalency and its function in regulating key developmental gene activation during early cell fate commitment.
Prof. Wei Xie from School of Life Science at Tsinghua University is the corresponding author of this work. Dr. Yunlong Xiang and Postdoc fellow Yu Zhang from CLS program at Tsinghua university are the co-first authors of this work. Prof. Lei Li from Institute of Zoology at Chinese Academy of Sciences and Prof. Sundeep Kalantry from Department of Human Genetics at University of Michigan also made important contributions to this study. This work was supported by the Beijing Municipal Science & Technology Commission, National Natural Science Foundation of China, Beijing Advanced Innovation Center for Structural Biology, National Basic Research Program of China, and the THU-PKU Center for Life Sciences. Prof. Wei Xie is also an HHMI International Research Scholar.
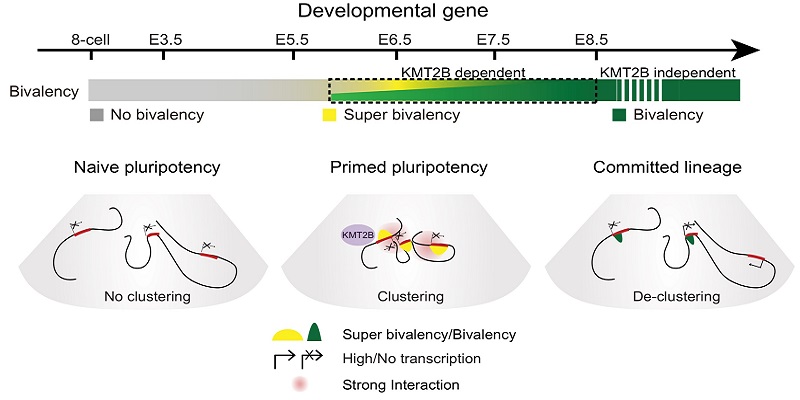
Dynamics of epigenome from na?ve pluripotency to committed lineages in vivo
